
Hardware, Software, Wetware, Co-Design Research
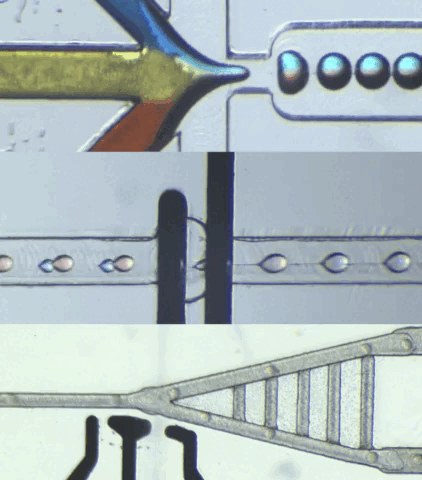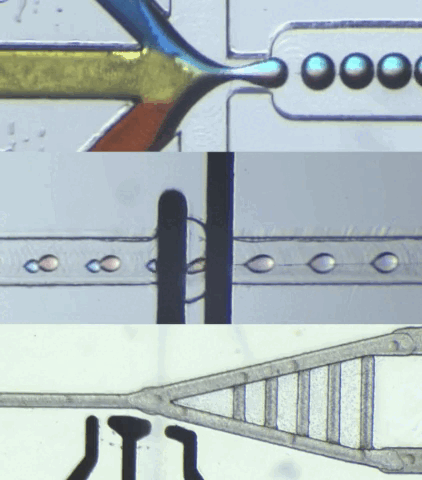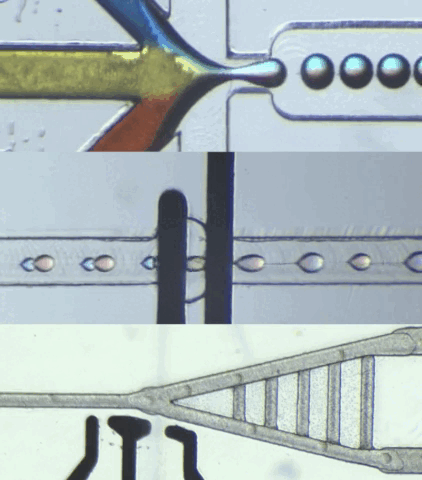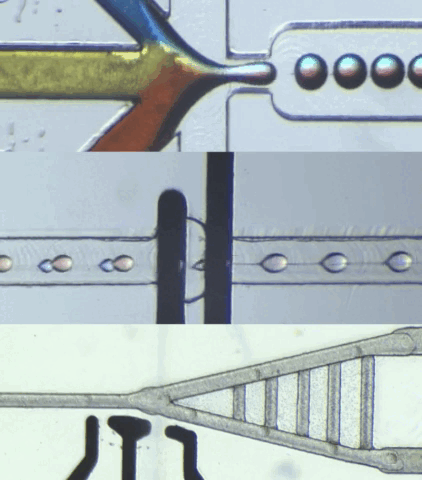
HARDWARE
CIDAR Lab is dedicated to developing custom biological-application hardware by converting high-level specifications into practical automated solutions, meeting requirements in terms of function, structure, and performance. We also specialize in building custom microfluidic hardware that control and process data for engineered biological systems, ensuring smooth integration with various applications.

SOFTWARE
CIDAR Lab creates software tools and algorithms to simplify the design, simulation, and analysis of engineered biological systems. Our software efficiently maps functional requirements to biological elements, encompassing applications like DNA compilation, cellular synthesis, and microfluidic device control. This automation enables researchers and engineers to innovate in the rapidly evolving fields of synthetic biology and bioinformatics.

WETWARE
Our research at CIDAR Lab focuses on engineering biological building blocks through well-defined performance parameters. As the foundation of our system designs, they are explicitly designed to be amendable to high throughput assembly protocols, and we store these elements both virtually and physically using online parts registries which provide convenient access for researchers working on bioengineered systems.
Design Automation
Software tools make synthetic biology more effective by integrating concepts from electronic design automation into the specification, design, and assembly of complex biological systems.
Interdisciplinary Approach
CIDAR is composed of a diverse team of undergraduates, graduate students, post-docs, and research staff from a range of programming, engineering and scientific backgrounds.
Mixed Purpose Workspaces
With both Computational and Wetlab facilities, CIDAR has workspaces for software development and biological engineering research across multiple departments at Boston University.
